Genomic Gymnastics in the Nucleocapsid Gene of SARS-CoV-2 During Transmission in Humans: Transposition of ACGAAC and Creation of the novel N2 Gene
William R. Gallaher
Mockingbird Nature Research Group, Pearl River, LA 70452, and
Emeritus Faculty. Dept. of Microbiology, Immunology and Parasitology, Louisiana State University, School of Medicine, New Orleans LA 70112
SUMMARY
Copy-choice transposition of the Transcriptional Regulatory Sequence (TRS) ACGAAC, to an area of partial sequence identity at the end of the Serine-Arginine (SR) rich region of the SARS-CoV-2 nucleocapsid, is likely to have created a novel canonical mRNA and gene specific for expression of the N2 dimerization domain of the nucleocapsid (N) protein of SARS-CoV-2. The transposition first occurred in the A2a lineage of the virus in China while circulating in the human population, but heretofore has only been recognized as the amino acid change 203RG204 to KR. While coronaviruses have a long history of such re-iterations of ACGAAC, this is the first time a virus has been observed to create a new canonical gene and mRNA during a human outbreak and while sequential sequencing documents its evolution. While its expression requires confirmation, the novel N2 gene may alter the dynamics of virion biogenesis.
INTRODUCTION
In their study of the structure and function of the SARS-CoV-2 nucleocapsid, first posted May 17, 2020, Ye et al (1) also briefly explored the genetic variability of the N gene within a large sampling of sequences obtained both in China before that date, as well as initial Genbank sequences from outside China. They found a mutational “hot spot” of variability at the end of the Serine-Arginine (SR)-rich region of N, centered around the last of the SR repeats. The most frequent genetic change they observed was what they termed a “triple-nucleotide substitution”, i.e. ggg to aac at 28881 to 28883 in the reference Hu-1 genome, that was out of frame and overlay two codons, changing 203RG204 to 203KR204. In the samples from China, this mutation was restricted to a fraction of the A2a genotype of lineage A, and comprised 15% of the total isolates. In the Genbank group, they constituted 10-15% of the total.
Virtually all of the details were contained in a supplementary figure S2 rather than in the more openly published main paper. What was evident in the sequence alignment in Figure S2A, but not expressed in the paper, was that the Genbank sequences containing the mutation were then of lineage B, and found as early as March 10, 2020 in Michigan.
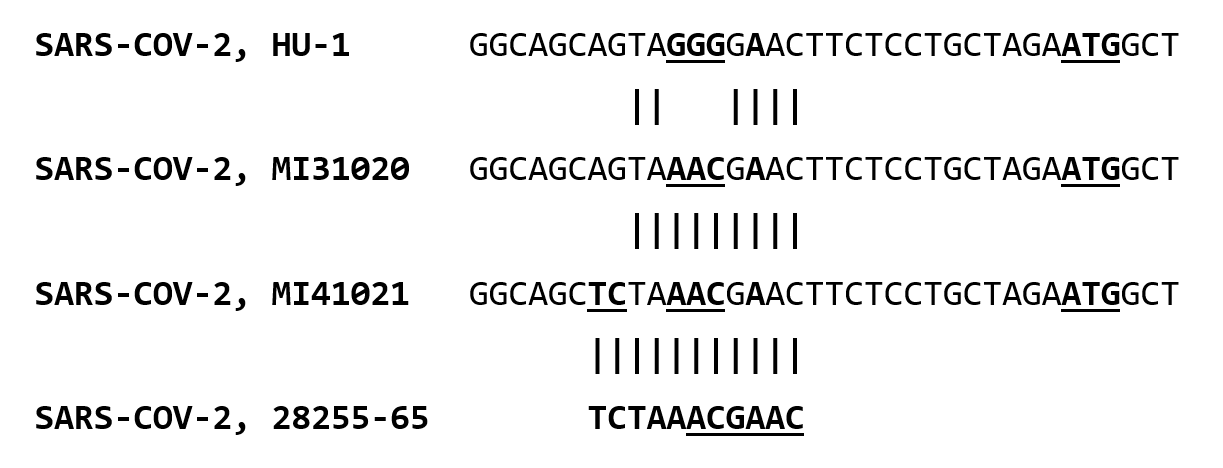
The alignment, adjusted slightly from that in Ye at al, for a reason to become apparent later, is shown in Figure 1:
Figure 1: Alignment of nucleotides 28853 through 28906 of SARS-CoV-2 reference sequence Hu-1 (Genbank NC_045512) corresponding to amino acids 195-211 of the nucleocapsid protein, with the aligning nucleotides and amino acids of Michigan isolate of SARS-CoV-2 from March 10, 2020 (Genbank MT439230)
The RG to KR change spread with the A2a lineage, and was actually first detected in the B lineage in western Europe and the United Kingdom, and then to the US. According to an evolutionary summary (2) first posted on Sep 24, 2020, it had by then spread to a number of foundational lineages throughout the world, comprising 25.7% of all then known sequences. These included A2a, B.1.1, the variant of concern B.1.1.7 (UK), B.1.1.10, B.1.1.14, B.1.1.25, B.1.1.26, the variant of concern B.1.1.28(Brazil), B.1.1.30 and B.1.1.33, again by Sep 24 of last year. The mutation is so widely spread that it is seldom included in a list of variant mutations. The change from RG to KR did not appear to confer any significant replicative advantage, but rather just make a basic region of a basic protein simply incrementally more basic.
While the literature on SARS-CoV-2 has become enormous and labyrinthine, it is difficult to find further mention, even in a parenthetical way. However, a good case can be made that this particular “triple-nucleotide substitution” is quite significant.
RESULTS
SIGNIFICANCE AT LEVEL OF PROTEIN
X-ray crystallography of the nucleocapsid protein shows a consistent picture, whether of SARS-CoV of 2003, MERS, or SARS-CoV-2 (1). The protein consists of four regions, an N-terminal globular RNA-binding domain (RnaBD) and a structurally independent C-terminal globular dimerization domain (DM), with the SR-rich region and a Joint (J) region in a “disordered” configuration in between (even though Ye et al was able to find some “residual helicity”).
However, the perspective of crystallography is limited, in that it often misses organization of what are conformationally dynamic regions of protein that only assume stable forms upon ligand interaction or environmental activation.
The symmetrical organization of the SR region is immediately obvious on inspection, as shown in Figures 2 through 5.
In Figure 2 is shown an amino acid alignment of the SR-rich region from SARS-CoV of 2003. The SARS-CoV-2 reference sequence Hu-1 of 2019, and the KR mutant sequence from the early (10 Mar) Michigan isolate. It should be noted that SARS-CoV-2 is not in any way directly derived from SARS-CoV of years earlier. Rather, they are distant relatives with a most recent common ancestor calculated by Boni et al (3) to have existed many centuries ago, around 1080AD. Nevertheless, the alignment shows an obviously symmetrical linear arrangement of SR dipeptides, i.e. (SR(X6)SR(X2)SRSR(X2)SR(X6)SR, conserved in each lineage for centuries. The RG to KR substitution disrupts this pattern at the carboxy-terminal end of the SR-rich region.
Figure 2: Alignment of the SR-rich regions of the nucleocapsids of SARS-CoV, Urbani of 2003 (Genbank AY278741), SARS-CoV-2, Hu-1 and SARS-CoV-2 Michigan isolate of Mar 10 2020, corresponding to amino acids 176 through 204 of Hu-1.
Figure 3 shows an extended beta projection of the Hu-1 sequence. It is immediately obvious that the arginine residues dominate the region, and that in this configuration they all lie on the same side of the extended peptide.
Figure 3: PyMol beta projection of the Hu-1 peptide from Figure 2 (www.pymol.org)
Figure 4 shows the same sequence now drawn as a helical wheel (4). The symmetry and sidedness of the region is striking, with all arginines clustered on one side of the helix while the other side is uniformly hydroxylated.
Figure 4: Projection of the Hu-1 peptide from Figure 2 on a helical wheel. Residues begin at the 0 degree position in the inner ring and amino acids after the first 18 project similarly in the outer ring.
Thus, whether the SR-rich region is in beta form or helical form, it displays a clear symmetry in the ancestral sequence of either the SARS-CoV or SARS-CoV-2 lineages.
Figure 5 shows the helical wheel projection of the KR mutant, that clearly distorts the ancestral symmetry.
Figure 5: Projection of the peptide from the SARS-CoV-2 mutant sequence from the Michigan isolate of March 10 2020, similarly on a helical wheel.
What positive effect that may have is uncertain, changing a symmetry conserved through the countless replication cycles since before King John signed the Magna Carta.
SIGNIFICANCE OF MUTATION AT THE LEVEL OF RNA
Indeed, the KR substitution in the nucleocapsid protein may be a cost paid for a much more significant change in the viral RNA sequence. Since the mutations were out-of-frame, it was easy to miss. An alignment of RNA sequences is shown in Figure 6, that focuses more tightly on the region of nt 28873 through 29906, and its similarity with the RNA sequence at the splice site within the orf8/N junction, about 600nt 5’ to the “trinucleotide substitution” mutation.
Figure 6: Alignment of nucleotides 28871 through 28906 of SARS-CoV-2, Hu-1 with the corresponding nucleotides of the KR mutant Michigan isolate of Mar 10, 2020 and mutant Michigan isolate of Apr 10, 2021, and the orf8/N junctional nucleotides of Hu-1, 28255 through 28265. Lines of identity are only shown with the orf8/N junctional site corresponding to the TRS splice site for the N gene.
It can be seen quite readily that the “triple nucleotide substitution” is instead a copy-choice transposition of the trinucleotide AAC derived from the orf8/N splice junction, that has the effect of duplicating the transcriptional regulatory signal (TRS), i.e. ACGAAC, to a new site in the genome at the end of the RNA encoding the SR-rich region. This site is already 66.7% ((6 out of 9) identical to the orf8/N splice junction in the SARS-CoV-2 Hu-1 reference strain, but the copy choice transposition renders it 100% identical over the nine critical bases comprising the splice junction site.
Further modification of the site is found a year later in viruses of the P1 Brazilian variant of concern, that is now widespread in the US. This further expansion of the RNA identity to 11/11 is accomplished by adding yet another two nucleotide substitutions that have escaped notice because they do not change amino acid 202 from serine, but instead substitutes an alternate codon for Serine, AGT to TCT. That this occurs by copy-choice transposition is obvious, since such a cluster of 5 mutations, 3 of them rarer transversions, is otherwise extremely unlikely. In particular, serine cannot be conserved at position 202 by any other path except by changing both of the first two nucleotides of the codon at once, and not sequentially.
Now, a suspicious mind might invoke PCR-mediated site-directed mutagenesis as a mechanism of change. However, these changes have occurred well after the introduction of the virus into the human population, and have evolved against quite different genetic backgrounds consistent with the time-dependent evolution of the virus, beginning in China but also occurring independently in several other places on earth, some quite remote. Furthermore, the virus itself is clearly capable of duplicating the ACGAAC sequence for the TRS splice acceptor site on its own. Since there are several such sites in a wide variety of coronavirus genomes, including several shared by both SARS-CoV and SARS-CoV-2, it is patently obvious that there is nothing about the duplication of this sequence by copy-choice transposition that is new. Indeed, it likely occurred many times in the evolution of coronaviruses long before humans walked the earth, let alone recognized that nucleic acid constituted the genetic material of life.
Finally, this transposition of the TRS is placed at a critical position in the N gene, prior to the entire sequence of the dimerization domain. An atg, in a moderately potent Kozak consensus context, lies just 13 nt 3’ to the transposition.
DISCUSSION
The effect of these mutations is very likely to create a novel mRNA and new open reading frame, beginning with Met 210, in the same frame as the whole nucleocapsid gene. In effect, the N2 gene.
The product of this novel N2 gene is predicted to extend for 210 amino acids, have a molecular weight of 23.1 kd, and an isoelectric point of 9.62. It should possess all of the properties of the independent dimerization domain of the nucleocapsid. Since nearly all studies of SARS-Cov-2 expression and molecular biology are performed on the reference strain for consistency (with the exception of the spike glycoprotein), something as simple as a gel profile of cells infected with a strain with this potentially new, independent gene has probably not been performed. Unless an immunoprecipitation was performed with N2-specific sera, it is unlikely that a new 23.1kd protein would have been noticed against the full protein profile in infected cells.
I no longer have a lab, so everyone should feel free to look.
While up to 33% of SARS-CoV-2 transcripts have been found to be “noncanonical” (5), without a vicinal 5’ TRS, these are not known to include initiation at Met210 of the N gene. However, there are some indications that a fragment of the nucleocapsid may be produced in an E. coli-based expression vector system (R.F. Garry, unpublished). Noncanonical messages are usually expressed at low levels, so even if initiation at Met210 occurs in the absence of the TRS, it would be expected to substantially increase when vicinal to a 5’ ACGAAC hexanucleotide.
Since this genetic event is common to both the B.1.1.7 and B.1.1.28 variants of concern, it has doubtlessly become extraordinarily widespread. It does not appear to be found in the B.1.617 India (“Delta”) variant that has rampaged through Southeast Asia, but in that sub-lineage 203R has been substituted with 203M, introducing a potential initiation codon at that site that may allow for noncanonical mRNAs to be produced with similar albeit lesser effect.
There are obviously no data to illuminate what effect an independently produced N2 protein would have on infection. However, one might speculate that, since virion biogenesis requires interaction of several cytoplasmic portions of the viral structural proteins S, E, M and N, an independent dimer of N2 might be able to initiate the process before the mature ribonucleoprotein complex arrived at the membrane. It might remain with the virion, or might indeed be displaced by the more mature and stable configuration complexed with RNA.
There are many examples of viral genes coding for more than one protein, or different versions of the same protein, and obviously also the duplication of genes or splice acceptor sites. I do not know of any such event that has occurred during transmission of a virus in the human population, while we have been indeed watching by intense sequencing of viral isolates. This is a novel and potentially significant event.
If the virus can do this, even multiple times, it can do just about anything by copy choice genetic gymnastics with respect to any of its many variable and hypervariable peptide regions. Given the previous constancy of the 202SR203 dipeptide, not even “constant” amino acids may be entirely excluded from change. With 371,000 new cases globally just yesterday, and just 75 days before the Fall/Winter season for respiratory disease arrives in the Northern Hemisphere, future prospects for the human race appear dim for the second winter of the pandemic
CONCLUSION:
A copy choice transposition, multiple times against different genetic backgrounds during human transmission of SARS-CoV-2, has duplicated the TRS hexanucleotide ACGAAC into a critical position of the nucleocapsid gene of SARS-CoV-2. With a potential initiation Met210 just 13 nt downstream, the transposition creates a molecular environment consistent with creation of a novel gene and mRNA that may independently express the carboxy-terminal dimerization domain of the nucleocapsid complex. This mutant configuration in globally widespread. While re-iteration of the TRS has doubtlessly occurred several previous times during coronavirus evolution, this is the first time such an event has been observed for any human virus during transmission in humans.
ACKNOWLEDGMENT: This communication is dedicated to the memory of my former wife, Betty Jean Burton Grier Gallaher RN who died Jan 10, my aunt, Connie A. Barrett, who died Feb 8, and my own dearly departed son, William Christopher Gallaher MS, who died May 19. But for COVID, they would likely still be with us and not have died within the span of 129 days in 2021. Requiescant in pace.
I thank my friend and collaborator of nearly 4 decades, Robert F. Garry, for helpful discussions, review of the manuscript, and access to his unpublished findings.
This work was produced with zero institutional or grant support, and zero conflicts of interest.
REFERENCES:
- Ye Q, West AMV, Silletti S, Corbett KD. Architecture and self-assembly of the SARS-CoV-2 nucleocapsid protein. Protein Sci. 2020 Sep;29(9):1890-1901. doi: 10.1002/pro.3909. Epub 2020 Aug 6. PMID: 32654247; PMCID: PMC7405475.Posted May 17 20
- Kumar S, Tao Q, Weaver S, Sanderford M, Caraballo-Ortiz MA, Sharma S, Pond SLK, Miura S. An evolutionary portrait of the progenitor SARS-CoV-2 and its dominant offshoots in COVID-19 pandemic. Mol Biol Evol. 2021 May 4:msab118. doi: 10.1093/molbev/msab118. Epub ahead of print. PMID: 33942847; PMCID: PMC8135569. Posted 92420
- EMBOSS: pepwheel (bioinformatics.nl)
- Boni MF, Lemey P, Jiang X, Lam TT, Perry BW, Castoe TA, Rambaut A, Robertson DL. Evolutionary origins of the SARS-CoV-2 sarbecovirus lineage responsible for the COVID-19 pandemic. Nat Microbiol. 2020 Nov;5(11):1408-1417. doi: 10.1038/s41564-020-0771-4. Epub 2020 Jul 28. PMID: 32724171.
- Nomburg J, Meyerson M, DeCaprio JA. Pervasive generation of non-canonical subgenomic RNAs by SARS-CoV-2. Genome Med. 2020 Dec 1;12(1):108. doi: 10.1186/s13073-020-00802-w. PMID: 33256807; PMCID: PMC7704119.