Genomic Evidence Links the Largest Dengue Outbreak in Senegal to Newly Circulating DENV-2 Lineage
Idrissa DIENG1,2*, Marie Henriette Dior Ndione1*, Mignane Ndiaye1,Bocar Sow3, Samba Niang Sagne3, Cheikh Talibouya Toure1, Sokhna Maï Diop1, Elisabeth Faye1, Agathe Shella Efire1, Ndeye Khady Diatou Ndiaye3, Al Ousseynou Seye1, Mouhamed Kane1,Mareme Seye Thiam3, Aboubacry Gaye3, Safietou Sankhe1,Babacar Diouf4, Alioune Gaye4, Cheikh Talla3, Ndongo Dia1, Diawo Diallo4, Ousmane Faye5, Mawlouth Diallo4, Cheikh Loucoubar3, Boubacar Diallo5, Abdourahmane Sow5, Mamadou Aliou Barry3, Moussa Moise Diagne1*, Gamou Fall1*, Oumar Faye1*
1Virology Department, Institut Pasteur de Dakar
2Animal Biology Department, Universite Cheikh Anta Diop de Dakar
3Epidemiology Clinical Reasearch and Data Science, Institut Pasteur de Dakar
4Zoology Medical, Institut Pasteur de Dakar
5Direction of Public Health, Institut Pasteur de Dakar
Correspondance: [email protected]
Abstract
In 2025, Senegal experienced an unprecedented nationwide dengue outbreak with 3633 confirmed cases (1)and characterized by co-circulation of DENV-1 and DENV-2, with DENV-2 as the predominant serotype. Molecular surveillance revealed persistence of DENV-1 lineage III_A.2, previously detected in 2024 and genetically close to strains from Mali and Nigeria, alongside the introduction of a new DENV-2 lineage (2II_F) closely related to strains from India and Réunion Island (2023–2024).
Clinically, severe dengue cases, including two fatalities, were mainly associated with DENV-2 infections. Genomic screening for antiviral resistance markers showed no mutations associated with Mosnodenvir resistance in either serotype, suggesting that circulating viruses remain susceptible. Altogether, these findings highlight the importance of ongoing syndromic arbovirus surveillance coupled with genomic data. They emphasize the urgent need for integrated genomic surveillance, antiviral preparedness, and vaccine performance reassessment to mitigate dengue burden in West Africa.
Introduction
Dengue virus (DENV), a mosquito-borne Orthoflavivirus with four antigenically distinct serotypes (DENV-1 to DENV- 4), represents a growing public health threat globally (2). However globally, recent years have shown increasing viral diversification and expansion (3), coinciding with climate variability (4) and increased regional mobility. These factors are known to fuel vector-borne disease global distribution and occurrence (5). Historically, Senegal has reported yearly dengue outbreaks since the late 2017 (6).
In 2024, the Senegalese dengue landscape was dominated by DENV-1 lineage III_A.2, genetically related to strains from Mali and Nigeria, indicating regional persistence (Unpublished Data). The 2025 season marked a major epidemiological shift, with the co-circulation of DENV-1 and DENV-2 across multiple administrative regions and an unprecedented number of clinical cases ever recorded since the implementation of the Sentinel Syndromic Surveillance system the 4S Network.
This study aimed to (i) characterize the molecular epidemiology of DENV-1 and DENV-2 during the 2025 outbreak in Senegal; (ii) infer the virus genotype, major and minor lineages and (iii) assess resistance mutations associated with the antiviral Mosnodenvir in order to evaluate its potential contribution to dengue burden mitigation strategies.
Materials and Methods
Syndromic Surveillance, Sample Collection and Laboratory Diagnosis
The first case of this current epidemic was recorded on July 22, 2025 in Kaolack region. It was a 45-year-old male without any travel history, who presented to the healthcare center with history of acute fever. To rule out arboviral etiology, 5ml blood sample was collected and shipped with the demographic and clinical form to Institut Pasteur de Dakar (IPD) for arbovirus testing. Results from molecular screening confirmed DENV infection.
From notification to this first suspected case through serum samples were collected from patients presenting with dengue-like illness in all regions of Senegal as part of the Syndromic Sentinel Surveillance of Senegal (4S network). Samples from participating sites were shipped to the Virology Department of the IPD for arbovirus testing. At IPD, arbovirus diagnostic was performed according to the protocol previously described by Dieng and colleagues (6). In case of DENV positivity by RT-qPCR the viral serotype was determined using multiplex RT-qPCR as previously described (7). Additionally, a subset of DENV+ samples with known viral serotype were subjected to whole genome sequencing and subsequently to phylogenetic analysis.
Sequencing and Phylogenetic Analysis
Twist Comprehensive Viral Research Panel (CVRP) hybrid capture followed by sequencing on an Illumina iSeq100 was used to generate whole-genome data from a subset of DENV-positive samples with cycle threshold (Ct) values ≤ 32, selected to cover the widest possible spatial distribution across Senegal. Library preparation, genomic sequencing, and processing of raw FASTQ reads to obtain viral consensus genomes were performed according to a previously described protocol (8). Raw data were analyzed using CZID platform (http://czid.org/; accessed on November 15 2025). Genotypes, Major lineage and Minor lineage of epidemic Senegalese sequences were assigned according using GenomeDetective (https://www.genomedetective.com/app/typingtool/virus/). Consensus sequences were aligned with reference global datasets maintained on Nexstrain (9) using MAFFT (10), and phylogenetic trees were inferred in IQ-TREE (11) under the GTR+F+I+G4 model with 1,000 ultrafast bootstraps. Temporal phylogenies were generated with TreeTime and visualized using ggtree (R package).
Mosnodenvir Resistance Screening using Drug platform
Amino acid positions previously associated with reduced sensitivity to Mosnodenvir were extracted from published experimental studies. Multiple sequence alignments were used to identify any deviations from the wild-type DENV-1 and DENV-2 references as previously described (12). Newly implemented tool namely DRUG was used to call amino acid changes on NS4B associated to Mosnodenvir resistance (https://idevfactory.shinyapps.io/DRUG/ ; Accessed on November 15, 2025).
Results
Epidemiological Overview
The 2025 dengue epidemic in Senegal recorded 3633 laboratory DENV confirmed samples among 15584 suspected cases at the date of December 12 2025. marking this highest annual number of dengue infections ever documented in the country since the implementation of the syndromic sentinel surveillance in 2015. Co-circulation of DENV-1 and DENV-2 was documented with DENV-2 accounting for approximately 98% of serotyped DENV+ samples (n = 789).
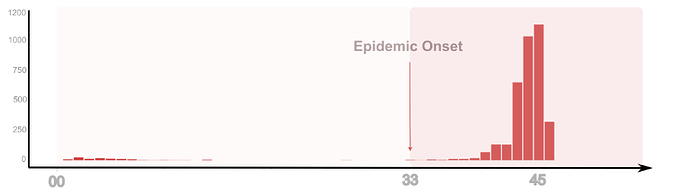
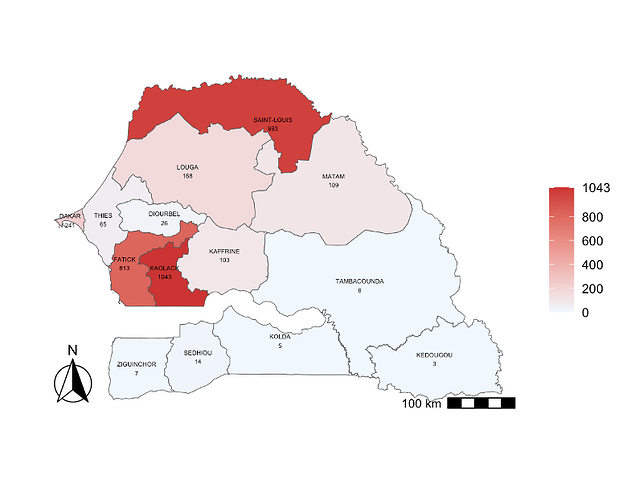
The outbreak displayed an early onset in week 33, peaking in week 45 (Figure 1). The majority of confirmed cases occurred in urban centers Kaolack (n = 1043), followed by Saint-Louis (n = 993), Fatick (n = 813). Exceptionally the DENV cases were recorded in Ziguinchor and Sedhiou the two areas exempt of confirmed acute dengue infection despite yearly dengue outbreak recorded since 2017 around the country (Figure 2). Two DENV infections related deaths occurred in two adult patients infected with DENV-2.
Phylogenetic Analysis
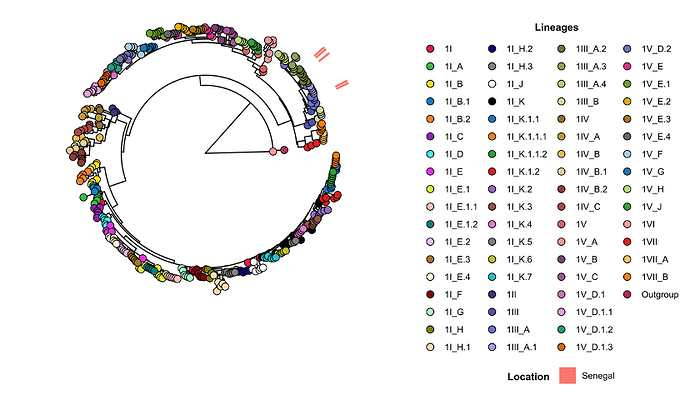
All DENV-1 sequences generated during this study (n=03) clustered within genotype III previously reported in Senegal and West Africa (6), specifically within major lineage 1III_A.2, which had been previously detected in 2021 in Senegal (13) (Figure 3). Phylogenetically, these strains were closely related to isolates from Mali (2023) and Nigeria (2022), suggesting ongoing endemic maintenance of this lineage within the subregion rather than recent reintroduction.
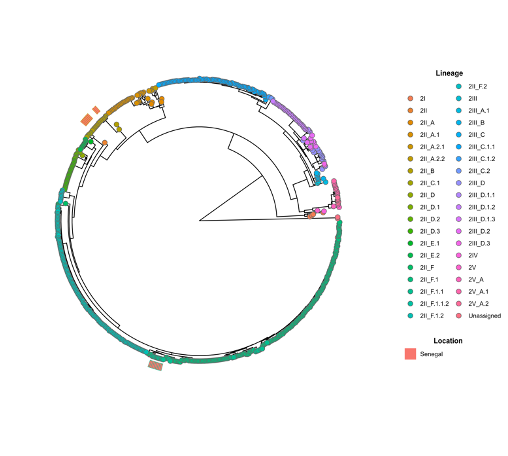
DENV-2 sequences (n = 11) belonged to genotype II namely cosmopolitan DENV-2 which is known to drive most of DENV-2 epidemic previously reported in West Africa (14), with all clustering within the new lineage 2II_F. This lineage showed strong phylogenetic affinity (bootstrap support >98%) with Indian and Réunion Island DENV-2 sequences from 2023–2024, indicating a likely importation from the southwestern Indian Ocean. Notably, these 2025 DENV-2 genomes showed clear separation from earlier Senegalese DENV-2 strains (6,15,16), confirming that the 2025 epidemic was driven by a newly introduced lineage rather than re-emergence of previously circulating DENV-2 variants (Figure 4).
Antiviral Resistance Analysis
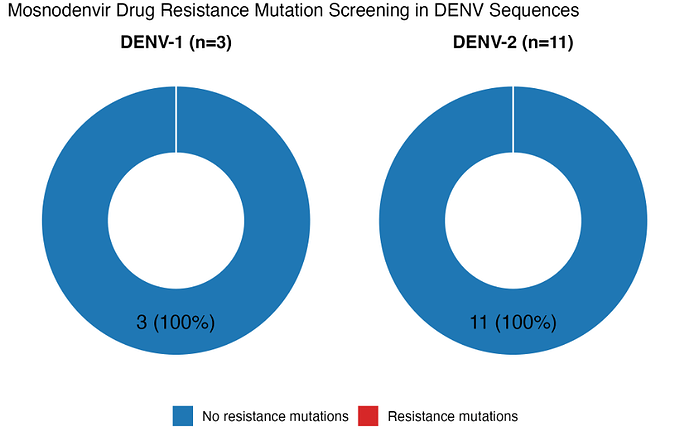
Alignment of the NS4B non-structural protein region of the NS4B revealed no mutations previously linked to Mosnodenvir resistance, such as substitutions at residues A91V, 94 (DENV-2 and 92 for DENV-1) and 108. Both DENV-1 and DENV-2 sequences retained the wild-type amino acid signatures at these positions (Figure 5), suggesting preserved sensitivity to the drug as previously described (12). These findings support the inclusion of Mosnodenvir in national dengue response strategies as a viable antiviral option should clinical use be warranted.
Discussion
The 2025 dengue epidemic in Senegal represents a turning point in the national dengue landscape, marked by the highest number of laboratory confirmed DENV cases since the implementation of the 4S network (6).
Additionally, it is marked by simultaneous circulation of DENV-1 and DENV-2 and the emergence of a new DENV-2 lineage (2II_F), phylogenetically close to viruses detected in India and Reunion Island in 2023-2024. The potential introduction of this lineage from the Indian Ocean region likely contributed to the unprecedented scale and severity of the outbreak. This corroborates previous studies that assessed that the introduction of a new virus variant in a partially immune population can drive large epidemics marked by an increased number of cases (17) and the occurrence of severe outcomes associated or not to fatal cases.
The maintenance of DENV-1 lineage III_A.2 reflects ongoing endemic persistence and regional connectivity with neighboring West African countries. Conversely, the emergence of DENV-2 lineage 2II_F, linked to severe cases and fatalities, signals a potential increase in virulence and the urgent need for enhanced genomic and clinical monitoring.
The absence of Mosnodenvir resistance mutations in both serotypes is an encouraging key finding. It indicates that circulating strains remain susceptible to this antiviral based on previously reported resistance associated mutations (12). Taken together, these findings support the potential integration of Mosnodenvir into therapeutic preparedness frameworks for dengue management in Senegal. Additionally, evaluation of the impact of this molecule on local African DENV strains should also be explored through in vitro experiment using Senegalese and African viral isolates.
Conclusion
This study provides comprehensive molecular and clinical insights into the 2025 dengue epidemic in Senegal, characterized by the following keys features:
-
Co-circulation of DENV-1 and DENV-2. With the reemergence of urban VDEN-2 after 05 year of none notification,
-
Predominance of a newly introduced DENV-2 Major lineage (2II_F) linked to a higher number of confirmed cases, a more severe disease picture and fatalities,
-
Local persistence of DENV-1 Major lineage 1III_A,
-
No evidence of Mosnodenvir resistance.
Together, these findings emphasize the importance of continuous sentinel syndromic surveillance of febrile illness to elucidate the contribution of dengue and related arbovirus, genomic surveillance to track shifts in viral lineages and serotype dominance, potential integration of antivirals like Mosnodenvir within dengue burden mitigation strategies, and reassessment of vaccine strategy and strain composition improve dengue control and prevention strategies within West Africa.
Figure 1. Kinetics of 2025 DENV virus outbreak in Senegal. From notification of the first case (2025 Dengue season) to the increased notification of confirmed cases. The barplot in red of confirmed DENV cases by PCR
Figure 2. Spatial distribution of detected DENV cases during the epidemic 2025. Confirmed DENV infection cases was detected in all administrative regions of the country.
Figure 3. Nextstrain tree of global DENV-1 strains colored by lineages. Senegalese sequences from previous years and from the ongoing 2025 outbreak belong to the same major lineages (1III_A). The 2025 epidemic is associated to strains clustering within the minor lineage DENV 1III_A.2 lineage.
Figure 4. Nextstrain tree of global DENV-2 strains colored by lineages. Senegalese sequences from previous years and from the ongoing 2025 outbreak belong two different major lineages 2II_B and 2II_F respectively. The 2025 epidemic is associated to strains clustering within DENV 2II_F lineage.
Figure 5. Mosnodenvir mutations associated resistance screening on circulating Senegalese VDEN 1-2 during 2025 epidemic. None of analyzed VDEN-1 / DENV-2 sequences harbor mutations related to Mosnodenvir resistance.
Acknowledgments
We thank the Senegalese Ministry of Health teams and the departments of the Institut Pasteur de Dakar for there contributions and technical support.
References
1. Bhatt S, Gething PW, Brady OJ, Messina JP, Farlow AW, Moyes CL, et al. The global distribution and burden of dengue. Nature. 25 avr 2013;496(7446):504‑7.
2. Haider N, Hasan MN, Onyango J, Asaduzzaman M. Global landmark: 2023 marks the worst year for dengue cases with millions infected and thousands of deaths reported. IJID Reg. déc 2024;13:100459.
3. Dieng I, Ndione MHD, Fall C, Diagne MM, Diop M, Gaye A, et al. Multifoci and multiserotypes circulation of dengue virus in Senegal between 2017 and 2018. BMC Infect Dis. 24 août 2021;21(1):867.
4. Nakase T, Giovanetti M, Obolski U, Lourenço J. Global transmission suitability maps for dengue virus transmitted by Aedes aegypti from 1981 to 2019. Sci Data. 12 mai 2023;10(1):275.
5. Acharya BK, Khanal L, Dhimal M. Increased thermal suitability elevates the risk of dengue transmission across the mid hills of Nepal. Harapan H, éditeur. PLOS One. 24 avr 2025;20(4):e0322031.
6. Dieng I, Talla C, Barry MA, Gaye A, Balde D, Ndiaye M, et al. The Spatiotemporal Distribution and Molecular Characterization of Circulating Dengue Virus Serotypes/Genotypes in Senegal from 2019 to 2023. Trop Med Infect Dis. 27 janv 2024;9(2):32.
7. Dieng I, Balde D, Talla C, Camara D, Barry MA, Sagne SN, et al. Molecular Evolution of Dengue Virus 3 in Senegal between 2009 and 2022: Dispersal Patterns and Implications for Prevention and Therapeutic Countermeasures. Vaccines. 28 sept 2023;11(10):1537.
8. Mourik K, Sidorov I, Carbo EC, Van Der Meer D, Boot A, Kroes ACM, et al. Comparison of the performance of two targeted metagenomic virus capture probe-based methods using reference control materials and clinical samples. St. George K, éditeur. J Clin Microbiol. 12 juin 2024;62(6):e00345-24.
9. Hadfield J, Megill C, Bell SM, Huddleston J, Potter B, Callender C, et al. Nextstrain: real-time tracking of pathogen evolution. Kelso J, éditeur. Bioinformatics. 1 déc 2018;34(23):4121‑3.
10. Katoh K, Misawa K, Kuma K ichi, Miyata T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 15 juill 2002;30(14):3059‑66.
11. Nguyen LT, Schmidt HA, von Haeseler A, Minh BQ. IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Mol Biol Evol. janv 2015;32(1):268‑74.
12. Bouzidi HS, Sen S, Piorkowski G, Pezzi L, Ayhan N, Fontaine A, et al. Genomic surveillance reveals a dengue 2 virus epidemic lineage with a marked decrease in sensitivity to Mosnodenvir. Nat Commun. 9 oct 2024;15(1):8667.
13. Dieng I, Barry MA, Talla C, Sow B, Faye O, Diagne MM, et al. Analysis of a Dengue Virus Outbreak in Rosso, Senegal 2021. Trop Med Infect Dis. 7 déc 2022;7(12):420.
14. Letizia AG, Pratt CB, Wiley MR, Fox AT, Mosore M, Agbodzi B, et al. Retrospective Genomic Characterization of a 2017 Dengue Virus Outbreak, Burkina Faso. Emerg Infect Dis [Internet]. juin 2022 [cité 15 sept 2022];28(6). Disponible sur: Retrospective Genomic Characterization of a 2017 Dengue Virus Outbreak, Burkina Faso - Volume 28, Number 6—June 2022 - Emerging Infectious Diseases journal - CDC
15. Dieng I, Ndiaye M, Ndione MH, Sankhe S, Diagne MM, Sagne SN, et al. Molecular Characterization of Circulating DENV-2 During Outbreak in Northern Senegal, Rosso 2018 [Internet]. LIFE SCIENCES; 2021 déc [cité 26 janv 2022]. Disponible sur: https://www.preprints.org/manuscript/202112.0270/v1
16. Dieng I, Diallo A, Ndiaye M, Mhamadi M, Diagne MM, Sankhe S, et al. Full genome analysis of circulating DENV‐2 in Senegal reveals a regional diversification into separate clades. J Med Virol. 5 août 2022;jmv.28027.
17. Cerpas C, Vásquez G, Moreira H, Juarez JG, Coloma J, Harris E, et al. Introduction of New Dengue Virus Lineages after COVID-19 Pandemic, Nicaragua, 2022. Emerg Infect Dis [Internet]. juin 2024 [cité 17 déc 2025];30(6). Disponible sur: Introduction of New Dengue Virus Lineages of Multiple Serotypes after COVID-19 Pandemic, Nicaragua, 2022 - Volume 30, Number 6—June 2024 - Emerging Infectious Diseases journal - CDC