Thank you for sharing! Had some free time and had a go at making some segment trees.
Brief methods:
- Downloaded all taxid 1980456 nucleotide sequences
- Separated by segment and made some preliminary alignments and fasttree phylogenies
- Manually picked the sequences clustering closest with the new virus and remade proper nucleotide alignments and trees for each segment (alignments: mafft localpair, trees: iqtree2, 10,000 UFbootstraps)
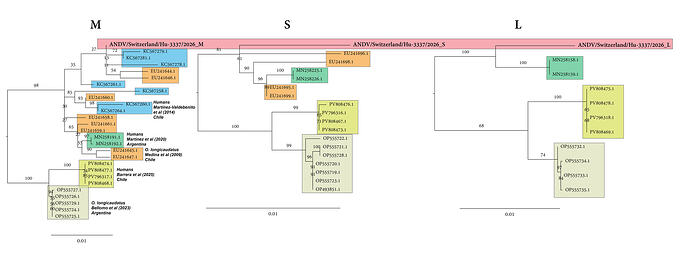
Here’s the trees (closest clade only) for all three segments:
Long story short, looks like a likely spillover from the O. longicaudatus rodent Argentina/Chile natural reservoir from a clade of Andes hantavirus that frequently infects humans.
I summarise the papers that describe the related viruses below:
KC accessions (blue):
Martinez-Valdebenito (2014): https://doi.org/10.3201/eid2010.140353
Human cases in Chile (Note: the sequences on this tree are the “control samples” described in the study and not the ones of the five patients of the outbreak described in the paper; M segment only).
EU accessions (orange):
Medina et al (2009): https://doi.org/10.1128/jvi.01057-08
Naturally infected Oligoryzomys longicaudatus (natural reservoir) from Chile
MN accessions (green):
Martinez et al (2020): https://doi.org/10.1056/NEJMoa2009040
2018-2019 outbreak in humans in Argentina
PV accessions (yellow):
Barrera et al (2025): https://doi.org/10.1016/j.crmicr.2025.100472
2024 outbreak in humans in Chile
OP accessions (grey):
Bellomo et al (2023): https://doi.org/10.1128/msphere.00018-23
Naturally infected Oligoryzomys longicaudatus (natural reservoir) from Argentina
All alignments, trees, and metadata here:
ANDV_summary_080526-spyros.zip (190.9 KB)
Let’s hope no more transmission of this one! ![]()
Spyros