Thank you everyone for the great work! Some additions and corrections on my previous tree after spending some time looking through previous papers and metadata.
Some of the KC and EU accessions on the M segment phylogeny corresponded to Gc and Gn sequences of the same virus isolate. I have now concatenated these into single sequences (removed KC567275.1-KC567264.1 due to potential sequencing errors) and trimmed the concatenated Gc/Gn coding sequence alignment to the parts with nucleotide available for all sequences.
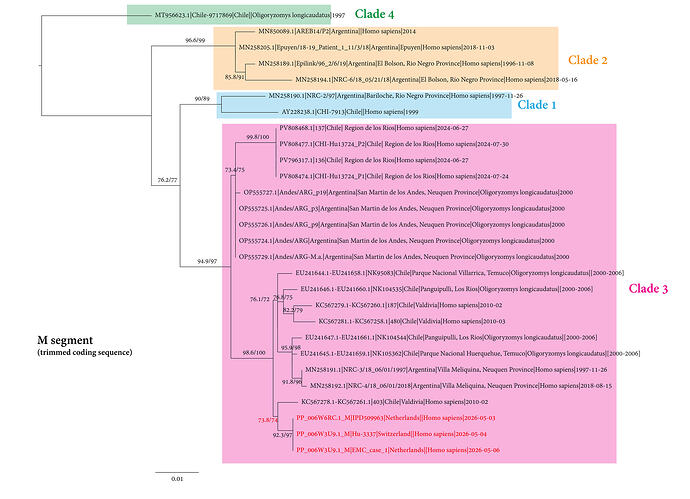
Along with the sequences closest to the new outbreak I have also included ANDV M segment sequences used in Bellomo et al. (2023) Fig2A (https://journals.asm.org/doi/10.1128/msphere.00018-23) and annotated ANDV clades in my tree accordingly:
Things to note:
- The single closest virus (based on this trimmed M segment alignment) is Andes virus isolate 403 published in Martinez-Valdebenito (2014, https://doi.org/10.3201/eid2010.140353) and sampled from a human patient in Chile in February 2010 but not associated with the 2011 outbreak described in the publication. Only the partial M segment used for this alignment is available for this virus.
- The closest complete virus sequences (with all three segments available) are isolates NRC-3 (MN258191.1) and NRC-4 (MN258192.1), also appearing as the closest in Emma’s nextstrain tree (Complete sequence of Orthohantavirus andesense virus: Swiss resident 2026 - #9 by emmahodcroft) and Andrew’s PearTree ( Experimental embedded trees for ANDV - #4 by arambaut ) are independent human isolates from Argentina published in Martinez et al (2020, https://doi.org/10.1056/NEJMoa2009040) but not associated with the super-spreader event described in the study. The 2018-2019 super-spreader event virus is from ANDV Clade 2, while the 2026 cruise ship virus is from ANDV Clade 3. Unclear what phenotypic implications this difference may have.
- Lastly there’s viruses sampled in rodents (Oligoryzomys longicaudatus) branching off across the tree suggesting that all human viruses with no documented association are likely independent spillovers from these rodents to humans.
Tree file, trimmed M segment alignment, and curated metadata here:
ANDV_M_segment-spyros-110526.zip (4.8 KB)
Feel free to use the above, Table S2 from Martinez et al is very useful for curating metadata: nejmoa2009040_appendix_1.pdf
Thanks again to all submitters and the Pathoplexus team for rapidly providing the data!