I got it. Thank you so much for your nice explanation and nice work!
I checked Saffold virus sequences.
The primer pair perfectly matches to Saffold virus sequences.
PSET results updated with three new 2019-nCoV genomes; 28 total genomes. Also adding three assays for comparison from the CDC (https://www.cdc.gov/coronavirus/2019-ncov/downloads/rt-pcr-panel-primer-probes.pdf). New genomes have resulted in mismatches in previously perfect alignments for some assays.
Table 1. Results from PSET analysis. The four Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Each assay was tested using 2019-nCoV (28 genomes) as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | provider | PT | TP | TN | PF | FP | FN |
|---|---|---|---|---|---|---|---|
| 2019-nCoV-noblis_1 | Noblis | 23 | 5 | 415 | NA | NA | NA |
| 2019-nCoV-noblis_2 | Noblis | 27 | 1 | 691 | NA | NA | NA |
| 2019-nCoV-noblis_3 | Noblis | 27 | 1 | 638 | NA | NA | NA |
| 2019-nCoV-noblis_4 | Noblis | 28 | NA | 356 | NA | 3 | NA |
| ncov_e_gene | Corman et al | 28 | NA | 75 | 353 | 15 | NA |
| ncov_n_gene | Corman et al | 27 | NA | 73 | NA | 339 | 1 |
| ncov_rdrp_1 | Corman et al | NA | 28 | 169 | 433 | 85 | NA |
| ncov_rdrp_2 | Corman et al | 1 | 27 | 586 | 1 | 61 | NA |
| 2019-nCoV_N1 | CDC | 27 | 1 | 381 | NA | NA | NA |
| 2019-nCoV_N2 | CDC | 27 | 1 | 371 | NA | NA | NA |
| 2019-nCoV_N3 | CDC | 28 | NA | 48 | NA | 346 | NA |
PSET results updated with new 2019-nCoV genomes; 46 total genomes.
Table 1. Results from PSET analysis. The four Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Each assay was tested using 2019-nCoV (46 genomes) as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | provider | PT | TP | TN | PF | FP | FN |
|---|---|---|---|---|---|---|---|
| 2019-nCoV-noblis_1 | Noblis | 36 | 10 | 415 | NA | NA | NA |
| 2019-nCoV-noblis_2 | Noblis | 45 | 1 | 689 | NA | NA | NA |
| 2019-nCoV-noblis_3 | Noblis | 45 | 1 | 637 | NA | NA | NA |
| 2019-nCoV-noblis_4 | Noblis | 46 | NA | 356 | NA | 3 | NA |
| ncov_e_gene | Corman et al | 46 | NA | 75 | 353 | 15 | NA |
| ncov_n_gene | Corman et al | 44 | 1 | 73 | NA | 339 | 1 |
| ncov_rdrp_1 | Corman et al | NA | 46 | 169 | 433 | 85 | NA |
| ncov_rdrp_2 | Corman et al | 1 | 45 | 586 | 1 | 61 | NA |
| cdc_n1 | CDC | 44 | 2 | 381 | NA | NA | NA |
| cdc_n2 | CDC | 45 | 1 | 371 | NA | NA | NA |
| cdc_n3 | CDC | 45 | 1 | 48 | NA | 346 | NA |
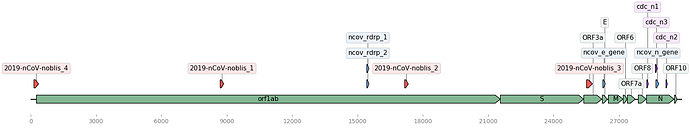
Figure 1. Updated genome map showing locations of all assays. Map of the Wuhan genome (NCBI Accession: MN908947.3) with assay signature locations (created using DNA Features Viewer Python library). Corman assays in blue, Noblis assays in red, CDC assays in purple, and gene regions in green.
PSET results updated with new 2019-nCoV genomes; 66 total genomes.
Table 1. Results from PSET analysis. The four Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Each assay was tested using 2019-nCoV (66 genomes) as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | provider | PT | TP | TN | PF | FP | FN |
|---|---|---|---|---|---|---|---|
| 2019-nCoV-noblis_1 | Noblis | 47 | 19 | 417 | NA | NA | NA |
| 2019-nCoV-noblis_2 | Noblis | 65 | 1 | 688 | NA | NA | NA |
| 2019-nCoV-noblis_3 | Noblis | 65 | 1 | 682 | NA | NA | NA |
| 2019-nCoV-noblis_4 | Noblis | 65 | NA | 357 | NA | 3 | 1 |
| ncov_e_gene | Corman et al | 66 | NA | 76 | 353 | 15 | NA |
| ncov_n_gene | Corman et al | 64 | 1 | 74 | NA | 339 | 1 |
| ncov_rdrp_1 | Corman et al | NA | 66 | 169 | 433 | 85 | NA |
| ncov_rdrp_2 | Corman et al | 1 | 65 | 586 | 1 | 61 | NA |
| cdc_n1 | CDC | 64 | 2 | 381 | NA | NA | NA |
| cdc_n2 | CDC | 65 | 1 | 371 | NA | NA | NA |
| cdc_n3 | CDC | 65 | 1 | 48 | NA | 346 | NA |
PSET results updated with new 2019-nCoV genomes; 96 total genomes. Now just using the subset of genomes on GISAID marked as high quality. Sequence IDs tested in this analysis listed here: ncov_ids_tested.zip (386 Bytes)
Assays continue to perform well against new genome submissions. Only one assay, 2019-nCoV-noblis_4, showing potential false negatives. Assay 2019-nCoV-noblis_1 has just one mismatch in the probe for 27 genomes, resulting in less perfect matches, but still likely functional for all 96 genomes.
Table 1. Results from PSET analysis. The four Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Each assay was tested using 2019-nCoV (96 genomes) as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | provider | PT | TP | TN | PF | FP | FN |
|---|---|---|---|---|---|---|---|
| 2019-nCoV-noblis_1 | Noblis | 69 | 27 | 372 | NA | NA | NA |
| 2019-nCoV-noblis_2 | Noblis | 96 | NA | 568 | NA | NA | NA |
| 2019-nCoV-noblis_3 | Noblis | 96 | NA | 439 | NA | NA | NA |
| 2019-nCoV-noblis_4 | Noblis | 94 | NA | 343 | NA | 3 | 2 |
| ncov_e_gene | Corman et al | 96 | NA | 42 | 353 | 15 | NA |
| ncov_n_gene | Corman et al | 95 | 1 | 55 | NA | 339 | NA |
| ncov_rdrp_1 | Corman et al | NA | 96 | 75 | 433 | 87 | NA |
| ncov_rdrp_2 | Corman et al | NA | 96 | 526 | 1 | 66 | NA |
| cdc_n1 | CDC | 95 | 1 | 363 | NA | NA | NA |
| cdc_n2 | CDC | 95 | 1 | 361 | NA | NA | NA |
| cdc_n3 | CDC | 94 | 2 | 17 | NA | 346 | NA |
PSET results updated with new 2019-nCoV genomes; 129 total genomes. Now just using the subset of genomes on GISAID marked as high quality. Sequence IDs tested in this analysis listed here: ncov_ids_tested.zip (437 Bytes)
Assays have more potential false negatives, although the majority of FNs (38 out of 42) are from samples collected from Pangolins. Only assays ncov_rdp_1 and cdc_n3 currently have no potential for FNs.
Table 1. Results from PSET analysis. The four Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Each assay was tested using 2019-nCoV (129 genomes) as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | provider | PT | TP | TN | PF | FP | FN |
|---|---|---|---|---|---|---|---|
| 2019-nCoV-noblis_1 | Noblis | 91 | 32 | 372 | NA | NA | 6 |
| 2019-nCoV-noblis_2 | Noblis | 123 | NA | 568 | NA | NA | 6 |
| 2019-nCoV-noblis_3 | Noblis | 122 | 2 | 439 | NA | NA | 5 |
| 2019-nCoV-noblis_4 | Noblis | 121 | 5 | 343 | NA | 3 | 3 |
| ncov_e_gene | Corman et al | 127 | 1 | 42 | 353 | 15 | 1 |
| ncov_n_gene | Corman et al | 122 | 2 | 55 | NA | 339 | 5 |
| ncov_rdrp_1 | Corman et al | NA | 129 | 75 | 433 | 87 | NA |
| ncov_rdrp_2 | Corman et al | NA | 124 | 526 | 1 | 66 | 5 |
| cdc_n1 | CDC | 122 | 2 | 363 | NA | NA | 5 |
| cdc_n2 | CDC | 122 | 1 | 361 | NA | NA | 6 |
| cdc_n3 | CDC | 121 | 8 | 17 | NA | 346 | NA |
PSET results updated with new 2019-nCoV genomes; 152 total genomes. Now just using the subset of genomes on GISAID marked as high quality. Sequence IDs tested in this analysis listed here: ncov_ids_tested.zip (475 Bytes)
Table 1. Results from PSET analysis. The four Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Each assay was tested using 2019-nCoV (152 genomes) as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | provider | PT | TP | TN | PF | FP | FN |
|---|---|---|---|---|---|---|---|
| 2019-nCoV-noblis_1 | Noblis | 106 | 39 | 373 | NA | NA | 7 |
| 2019-nCoV-noblis_2 | Noblis | 145 | NA | 568 | NA | NA | 7 |
| 2019-nCoV-noblis_3 | Noblis | 144 | 2 | 439 | NA | NA | 6 |
| 2019-nCoV-noblis_4 | Noblis | 144 | 5 | 343 | NA | 3 | 3 |
| ncov_e_gene | Corman et al | 149 | 2 | 42 | 353 | 15 | 1 |
| ncov_n_gene | Corman et al | 144 | 3 | 55 | NA | 339 | 5 |
| ncov_rdrp_1 | Corman et al | NA | 152 | 75 | 433 | 87 | NA |
| ncov_rdrp_2 | Corman et al | NA | 147 | 526 | 1 | 66 | 5 |
| cdc_n1 | CDC | 144 | 3 | 363 | NA | NA | 5 |
| cdc_n2 | CDC | 144 | 1 | 361 | NA | NA | 7 |
| cdc_n3 | CDC | 143 | 9 | 17 | NA | 346 | NA |
PSET results updated with new 2019-nCoV genomes; 190 total genomes. Now just using the subset of genomes on GISAID marked as high quality sampled from humans (pangolin samples removed). Sequence IDs tested in this analysis listed here: ncov_ids_tested.zip (534 Bytes)
Table 1. Results from PSET analysis. The four Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Each assay was tested using 2019-nCoV (190 genomes) as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | provider | PT | TP | TN | PF | FP | FN |
|---|---|---|---|---|---|---|---|
| 2019-nCoV-noblis_1 | Noblis | 134 | 56 | 373 | 0 | 0 | 0 |
| 2019-nCoV-noblis_2 | Noblis | 188 | 2 | 568 | 0 | 0 | 0 |
| 2019-nCoV-noblis_3 | Noblis | 189 | 1 | 439 | 0 | 0 | 0 |
| 2019-nCoV-noblis_4 | Noblis | 184 | 0 | 343 | 0 | 3 | 6 |
| ncov_e_gene | Corman et al | 187 | 2 | 42 | 353 | 15 | 1 |
| ncov_n_gene | Corman et al | 189 | 1 | 55 | 0 | 339 | 0 |
| ncov_rdrp_1 | Corman et al | 190 | 0 | 75 | 433 | 87 | 0 |
| ncov_rdrp_2 | Corman et al | 190 | 0 | 526 | 1 | 66 | 0 |
| cdc_n1 | CDC | 188 | 2 | 363 | 0 | 0 | 0 |
| cdc_n2 | CDC | 189 | 1 | 361 | 0 | 0 | 0 |
| cdc_n3 | CDC | 183 | 7 | 17 | 0 | 346 | 0 |
PSET results updated with new 2019-nCoV genomes; 432 total genomes. Just using the subset of genomes on GISAID marked as high quality sampled from humans. Sequence IDs tested in this analysis listed here: 432_ids.zip (833 Bytes)
Noblis assay 4 showing some potential FNs, but most of these appear to be the result of gaps or sequencing errors for some sequences at the very start of the genome. This assay falls within the first 500bp of the genome. All other assays still performing very well in silico against new sequences.
Table 1. Results from PSET analysis. The four Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Each assay was tested using 2019-nCoV (432 genomes) as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | provider | PT | TP | TN | PF | FP | FN |
|---|---|---|---|---|---|---|---|
| 2019-nCoV-noblis_1 | Noblis | 325 | 105 | 373 | 0 | 0 | 2 |
| 2019-nCoV-noblis_2 | Noblis | 422 | 10 | 568 | 0 | 0 | 0 |
| 2019-nCoV-noblis_3 | Noblis | 431 | 1 | 439 | 0 | 0 | 0 |
| 2019-nCoV-noblis_4 | Noblis | 409 | 4 | 343 | 0 | 3 | 19 |
| ncov_e_gene | Corman et al | 428 | 3 | 42 | 353 | 15 | 1 |
| ncov_n_gene | Corman et al | 431 | 1 | 55 | 0 | 339 | 0 |
| ncov_rdrp_1 | Corman et al | 0 | 432 | 75 | 433 | 87 | 0 |
| ncov_rdrp_2 | Corman et al | 0 | 432 | 526 | 1 | 66 | 0 |
| cdc_n1 | CDC | 429 | 3 | 363 | 0 | 0 | 0 |
| cdc_n2 | CDC | 430 | 2 | 361 | 0 | 0 | 0 |
| cdc_n3 | CDC | 412 | 20 | 17 | 0 | 346 | 0 |
PSET results updated with new 2019-nCoV genomes; 655 total genomes. Just using the subset of genomes on GISAID marked as high quality sampled from humans. Sequence IDs tested in this analysis listed here: 655_ids.zip (1.4 KB)
Previous Noblis assays showed some false negatives due to Ns, sequence gaps, and in one case the assay’s position at the very start of the genome. These have been replaced with five new assays generated at a later date using 96 complete genomes. The Noblis.57 assay and the German ncov_e_gene assay each have one FN that’s due to a stretch of Ns. All other assays still performing very well in silico against new sequences.
Table 1. Results from PSET analysis. The five Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Each assay was tested using 2019-nCoV (655 genomes) as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | provider | PT | TP | TN | PF | FP | FN |
|---|---|---|---|---|---|---|---|
| Noblis.12 | Noblis | 653 | 2 | 2 | 0 | 0 | 0 |
| Noblis.40 | Noblis | 635 | 20 | 316 | 0 | 0 | 0 |
| Noblis.42 | Noblis | 655 | 0 | 275 | 0 | 0 | 0 |
| Noblis.44 | Noblis | 655 | 0 | 277 | 0 | 0 | 0 |
| Noblis.57 | Noblis | 654 | 0 | 359 | 0 | 0 | 1 |
| ncov_e_gene | Corman et al | 651 | 3 | 42 | 353 | 15 | 1 |
| ncov_n_gene | Corman et al | 652 | 3 | 55 | 0 | 339 | 0 |
| ncov_rdrp_1 | Corman et al | 0 | 655 | 75 | 433 | 87 | 0 |
| ncov_rdrp_2 | Corman et al | 1 | 654 | 526 | 1 | 66 | 0 |
| cdc_n1 | CDC | 647 | 8 | 363 | 0 | 0 | 0 |
| cdc_n2 | CDC | 653 | 2 | 361 | 0 | 0 | 0 |
| cdc_n3 | CDC | 628 | 27 | 17 | 0 | 346 | 0 |
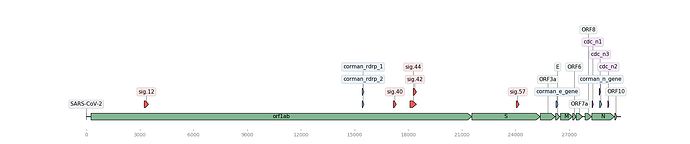
Figure 1. Map of the SARS-CoV-2 genome (NCBI Accession: MN908947.3) with assay signature locations (created using DNA Features Viewer Python library). Noblis assays in red, Corman assays in blue, CDC assays in purple, and gene regions in green.
PSET results updated with new 2019-nCoV genomes; 1620 total genomes. Just using the subset of genomes on GISAID marked as high quality sampled from humans. Sequence IDs tested in this analysis listed here: 1620_ids.zip (12.2 KB)
The Noblis.57 assay and the German ncov_e_gene assay still each have one FN that’s due to a stretch of Ns. All other assays still performing very well in silico against new sequences.
Table 1. Results from PSET analysis. The five Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Each assay was tested using 2019-nCoV (1620 genomes) as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | provider | PT | TP | TN | PF | FP | FN |
|---|---|---|---|---|---|---|---|
| Noblis.12 | Noblis | 1616 | 4 | 2 | 0 | 0 | 0 |
| Noblis.40 | Noblis | 1546 | 74 | 316 | 0 | 0 | 0 |
| Noblis.42 | Noblis | 1620 | 0 | 275 | 0 | 0 | 0 |
| Noblis.44 | Noblis | 1619 | 1 | 277 | 0 | 0 | 0 |
| Noblis.57 | Noblis | 1605 | 14 | 359 | 0 | 0 | 1 |
| ncov_e_gene | Corman et al | 1616 | 3 | 42 | 353 | 15 | 1 |
| ncov_n_gene | Corman et al | 1608 | 12 | 55 | 0 | 339 | 0 |
| ncov_rdrp_1 | Corman et al | 0 | 1620 | 75 | 433 | 87 | 0 |
| ncov_rdrp_2 | Corman et al | 1 | 1619 | 526 | 1 | 66 | 0 |
| cdc_n1 | CDC | 1591 | 29 | 363 | 0 | 0 | 0 |
| cdc_n2 | CDC | 1616 | 4 | 361 | 0 | 0 | 0 |
| cdc_n3 | CDC | 1560 | 60 | 17 | 0 | 346 | 0 |
Dear S. Sozhamannan,
please can you share the new primer and probe sequences of Noblis.12-57 as you did with 2019-nCoV-noblis_1-4? That would be great.
BTW: Do you know the primer sequences used by doi:10.1128/JCM.00310-20? They are not provided with their paper and the author did not respond within the last 12 days.
Thanks!
Sure thing! Here they are below. The full amplicon sequences are provided with the primers in brackets and probes in parentheses. We do not know the primers sequences used in that paper, but if we happen to find them we will let you know.
| Identifier | Amplicon |
|---|---|
| Noblis.12 | [ACGGCAGTGAGGACAATCAG]ACAACTACTATTCAAACAATTGTTGAGGTTCAACCTCAATTAGAGATGGAACTTACACCAGTTGTTCAGACTATTGAAGTGAATAGTTTTAGTGGTTATTTAAAACTTACTGACAATGTATACATTAAAAATGCAGACATTGTGGAAGAAGCTAAAAAGGTAAAA(CCAACAGTGGTTGTTAATGCAGCCA)ATGTTTACCTTAA[ACATGGAGGAGGTGTTGCAG] |
| Noblis.40 | [GCCGCTGTTGATGCACTATG]TGAGAAGGCATTAAAATATTTGCCTATAGATAAATGTAGTAGAATTATACCTGC(ACGTGCTCGTGTAGAGTGTTTTGAT)AAATTCAAAGTGAATTCAACATTAGAACAGTATGTCTTTTGTACTGTAA[ATGCATTGCCTGAGACGACA] |
| Noblis.42 | [TGTACGTGCATGGATTGGCT](TCGATGTCGAGGGGTGTCATGCT)ACTAGAGAAGCTGTTGGTACCAATTTACCTTTACAGCTAGGTTTTTCTACAGGTGTTAACCTAGTTGCTGTACCTACAGGTTATGTTGATACACCTAATAATACAGATTTTTCCAGAGT[TAGTGCTAAACCACCGCCTG] |
| Noblis.44 | [CAGGCACCTACACACCTCAG]TGTTGACACTAAATTCAAAACTGAAGGTTTATGTGTTGACATACCTGGCATACCTAAGGACATGACCTATAGAAGACTCATCTCTATGATGGGTTTTAAAATGAATTATCAAGTTAATGGTTACCCTAACATGTTTATCACCCGCGAAGAAGCTATAAGACATGTACGTGCATGGAT(TGGCTTCGATGTCGAGGGGTGT)CATGCTACTAGAGAAGCTGTTGGTACCAATTTACCTTTACAGCTAGGTTTTTCTACAGGTGTTAACCTAGTTGCTGTACCTACAGGTTATGTTGATACACCTAATAATACAGATTTTTCCAGAGT[TAGTGCTAAACCACCGCCTG] |
| Noblis.57 | [TGCAGATGCTGGCTTCATCA]AACAATATGGTGATTGCCTTGGTGATATTGCTGCTAGAGACCTCATTTGTGCACAAAAGTTTA(ACGGCCTTACTGTTTTGCCACCT)TTGCTCACAGATGAAATGATTGCTCAATACACTT[CTGCACTGTTAGCGGGTACA] |
**Small error identified in the results Table 1 from previous post on 4/10. Fixed below. (4/15/2020)
PSET results updated with new 2019-nCoV genomes; 3996 total genomes. Just using the subset of genomes on GISAID marked as high quality sampled from humans. Sequence IDs tested in this analysis listed here: 3996_ids.zip (29.1 KB)
Table 1. Results from PSET analysis. The five Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Each assay was tested using 2019-nCoV (3996 genomes) as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | provider | PT | TP | TN | PF | FP | FN |
|---|---|---|---|---|---|---|---|
| Noblis.12 | Noblis | 3984 | 12 | 2 | 0 | 0 | 0 |
| Noblis.40 | Noblis | 3855 | 140 | 316 | 0 | 0 | 1 |
| Noblis.42 | Noblis | 3990 | 6 | 275 | 0 | 0 | 0 |
| Noblis.44 | Noblis | 3988 | 8 | 277 | 0 | 0 | 0 |
| Noblis.57 | Noblis | 3967 | 25 | 359 | 0 | 0 | 4 |
| ncov_e_gene | Corman et al | 3983 | 7 | 42 | 353 | 15 | 6 |
| ncov_n_gene | Corman et al | 3965 | 31 | 55 | 0 | 339 | 0 |
| ncov_rdrp_1 | Corman et al | 0 | 3995 | 75 | 433 | 87 | 1 |
| ncov_rdrp_2 | Corman et al | 1 | 3995 | 526 | 1 | 66 | 0 |
| cdc_n1 | CDC | 3919 | 77 | 363 | 0 | 0 | 0 |
| cdc_n2 | CDC | 3982 | 14 | 361 | 0 | 0 | 0 |
| cdc_n3 | CDC | 3883 | 113 | 17 | 0 | 346 | 0 |
Table 2. False negative hits along with affected assay and sequence identifiers.
| num | identifier | accession | outcome |
|---|---|---|---|
| 1 | ncov_e_gene | France_ARA10968_2020_EPI_ISL_418431_2020-03-18 | FN |
| 2 | ncov_e_gene | USA_NY-NYUMC33_2020_EPI_ISL_418979_2020-03-17 | FN |
| 3 | ncov_e_gene | Italy_TE4959_2020_EPI_ISL_418259_2020-03-14 | FN |
| 4 | ncov_e_gene | France_ARA12626_2020_EPI_ISL_420612_2020-03-23 | FN |
| 5 | ncov_e_gene | France_ARA13074_2020_EPI_ISL_420622_2020-03-24 | FN |
| 6 | ncov_e_gene | France_Lyon_06042_2020_EPI_ISL_417333_2020-03-04 | FN |
| 7 | ncov_rdrp_1 | Australia_QLDID922_2020_EPI_ISL_418799_2020-02-28 | FN |
| 8 | Noblis.40 | USA_WA-S7_2020_EPI_ISL_416462_2020-02-24 | FN |
| 9 | Noblis.57 | Beijing_IVDC-BJ-005_2020_EPI_ISL_408485_2020-01-18 | FN |
| 10 | Noblis.57 | Canada_ON_PHL2223_2020_EPI_ISL_418381_2020 | FN |
| 11 | Noblis.57 | Canada_ON_PHL2273_2020_EPI_ISL_418383_2020 | FN |
| 12 | Noblis.57 | Canada_ON_PHL6884_2020_EPI_ISL_418343_2020-03-10 | FN |
Big Update-- Added 14 new assays to monitor. Updated NCBI databases (nt, env_nt) – 4/16/2020. Updated to 6,835 COVID-19 WGS from GISAID. Sequence IDs tested in this analysis listed here: 6835_ids.zip (44.5 KB)
Table 1. Results from PSET analysis. The five Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Along with new assays from China, Hong Kong, Thailand, Japan, and France. Each assay was tested using 2019-nCoV (6,835 genomes) as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | type | PT | TP | FN | Percent True | PF | FP | TN |
|---|---|---|---|---|---|---|---|---|
| Noblis.12 | Probe | 6793 | 41 | 1 | 99.99% | 0 | 0 | 4 |
| Noblis.40 | Probe | 6524 | 310 | 1 | 99.99% | 0 | 1 | 318 |
| Noblis.42 | Probe | 6825 | 8 | 2 | 99.97% | 0 | 1 | 0 |
| Noblis.44 | Probe | 6823 | 11 | 1 | 99.99% | 0 | 1 | 31 |
| Noblis.57 | Probe | 6779 | 50 | 6 | 99.91% | 0 | 1 | 7 |
| ncov_e_gene | Probe | 6813 | 13 | 9 | 99.87% | 361 | 15 | 3 |
| ncov_n_gene | Probe | 6784 | 51 | 0 | 100.00% | 0 | 340 | 27 |
| ncov_rdrp_1 | Probe | 0 | 6834 | 1 | 99.99% | 430 | 59 | 239 |
| ncov_rdrp_2 | Probe | 1 | 6834 | 0 | 100.00% | 2 | 37 | 677 |
| cdc_n1 | Probe | 6689 | 146 | 0 | 100.00% | 0 | 2 | 0 |
| cdc_n2 | Probe | 6787 | 26 | 22 | 99.68% | 0 | 1 | 330 |
| cdc_n3 | Probe | 6683 | 145 | 7 | 99.90% | 1 | 3 | 1 |
| China_N | Probe | 5606 | 47 | 1182 | 82.71% | 0 | 2 | 2 |
| China_ORF1ab | Probe | 6753 | 63 | 19 | 99.72% | 1 | 0 | 348 |
| France_nCoV_IP2 | Probe | 6820 | 14 | 1 | 99.99% | 1 | 0 | 1 |
| France_nCoV_IP4 | Probe | 6816 | 19 | 0 | 100.00% | 0 | 0 | 3 |
| HKU-N | Probe | 6762 | 48 | 25 | 99.63% | 320 | 9 | 4 |
| HKU-ORF1b-nsp14 | Probe | 6807 | 27 | 1 | 99.99% | 1 | 0 | 0 |
| Japan_NIID_2019-nCOV_N | Probe | 0 | 6812 | 23 | 99.66% | 0 | 0 | 360 |
| Japan_NIID_WH-1_F24381 | Outer | 6777 | 58 | 0 | 100.00% | 0 | 1 | 455 |
| Japan_NIID_WH-1_F501 | Outer | 6723 | 100 | 12 | 99.82% | 0 | 1 | 3 |
| Japan_NIID_WH-1_F509 | Outer | 6769 | 52 | 14 | 99.80% | 1 | 0 | 3 |
| Japan_NIID_WH-1_Seq_F24383 | Outer | 6778 | 57 | 0 | 100.00% | 0 | 0 | 456 |
| Japan_NIID_WH-1_Seq_F519 | Outer | 6795 | 22 | 18 | 99.74% | 0 | 1 | 3 |
| Japan_WuhanCoV-spk1 | Outer | 6786 | 49 | 0 | 100.00% | 1 | 1 | 454 |
| Thailand_WH-NIC_N | Probe | 6760 | 75 | 0 | 100.00% | 0 | 2 | 0 |
Updated to 7,903 COVID-19 WGS from GISAID. Just using the subset of genomes on GISAID marked as high quality sampled from humans.
Table 1. Results from PSET analysis. The five Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Along with new assays from China, Hong Kong, Thailand, Japan, and France. Each assay was tested using 2019-nCoV (7,903 genomes) as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | type | PT | TP | FN | Percent True | PF | FP | TN |
|---|---|---|---|---|---|---|---|---|
| Noblis.12 | Probe | 7859 | 43 | 1 | 99.99% | 0 | 0 | 680 |
| Noblis.40 | Probe | 7569 | 333 | 1 | 99.99% | 0 | 1 | 1420 |
| Noblis.42 | Probe | 7892 | 9 | 2 | 99.97% | 0 | 2 | 1136 |
| Noblis.44 | Probe | 7890 | 12 | 1 | 99.99% | 0 | 1 | 1786 |
| Noblis.57 | Probe | 7837 | 59 | 7 | 99.91% | 0 | 1 | 853 |
| ncov_e_gene | Probe | 7878 | 16 | 9 | 99.89% | 361 | 15 | 53 |
| ncov_n_gene | Probe | 7841 | 60 | 2 | 99.97% | 0 | 340 | 72 |
| ncov_rdrp_1 | Probe | 0 | 7902 | 1 | 99.99% | 433 | 105 | 5967 |
| ncov_rdrp_2 | Probe | 1 | 7902 | 0 | 100.00% | 2 | 73 | 5916 |
| cdc_n1 | Probe | 7681 | 221 | 1 | 99.99% | 0 | 2 | 381 |
| cdc_n2 | Probe | 7848 | 32 | 23 | 99.71% | 0 | 1 | 372 |
| cdc_n3 | Probe | 7725 | 176 | 2 | 99.97% | 1 | 353 | 36 |
| China_N | Probe | 6514 | 58 | 1331 | 83.16% | 0 | 2 | 691 |
| China_ORF1ab | Probe | 7819 | 65 | 19 | 99.76% | 1 | 0 | 367 |
| France_nCoV_IP2 | Probe | 7866 | 36 | 1 | 99.99% | 1 | 0 | 367 |
| France_nCoV_IP4 | Probe | 7877 | 25 | 1 | 99.99% | 0 | 0 | 1190 |
| HKU-N | Probe | 7812 | 65 | 26 | 99.67% | 327 | 33 | 34 |
| HKU-ORF1b-nsp14 | Probe | 7872 | 30 | 1 | 99.99% | 324 | 29 | 452 |
| Japan_NIID_2019-nCOV_N | Probe | 0 | 7879 | 24 | 99.70% | 0 | 0 | 384 |
| Japan_NIID_WH-1_F24381 | Outer | 7836 | 67 | 0 | 100.00% | 0 | 1 | 601 |
| Japan_NIID_WH-1_F501 | Outer | 7762 | 128 | 13 | 99.84% | 0 | 1 | 370 |
| Japan_NIID_WH-1_F509 | Outer | 7826 | 62 | 15 | 99.81% | 1 | 0 | 369 |
| Japan_NIID_WH-1_Seq_F24383 | Outer | 7838 | 65 | 0 | 100.00% | 0 | 0 | 606 |
| Japan_NIID_WH-1_Seq_F519 | Outer | 7852 | 29 | 22 | 99.72% | 0 | 1 | 369 |
| Japan_WuhanCoV-spk1 | Outer | 7845 | 58 | 0 | 100.00% | 1 | 1 | 594 |
| Thailand_WH-NIC_N | Probe | 7822 | 80 | 1 | 99.99% | 0 | 2 | 375 |
Updated to 15,411 COVID-19 WGS from GISAID. Using all genomes sampled from humans regardless of quality.
Table 1. Results from PSET analysis. The five Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Along with assays from China, Hong Kong, Thailand, Japan, and France. Each assay was tested using 2019-nCoV (15,411 genomes) as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | type | PT | TP | FN | Percent True | PF | FP | TN |
|---|---|---|---|---|---|---|---|---|
| Noblis.12 | Probe | 15251 | 71 | 89 | 99.42% | 0 | 0 | 680 |
| Noblis.40 | Probe | 14742 | 606 | 63 | 99.59% | 0 | 1 | 1420 |
| Noblis.42 | Probe | 15327 | 23 | 61 | 99.60% | 0 | 2 | 1136 |
| Noblis.44 | Probe | 15130 | 36 | 245 | 98.41% | 0 | 1 | 1786 |
| Noblis.57 | Probe | 15165 | 117 | 129 | 99.16% | 0 | 1 | 853 |
| ncov_e_gene | Probe | 15278 | 48 | 85 | 99.45% | 361 | 15 | 53 |
| ncov_n_gene | Probe | 15222 | 116 | 73 | 99.53% | 0 | 340 | 72 |
| ncov_rdrp_1 | Probe | 0 | 15366 | 45 | 99.71% | 433 | 105 | 5967 |
| ncov_rdrp_2 | Probe | 1 | 15369 | 41 | 99.73% | 2 | 73 | 5916 |
| cdc_n1 | Probe | 15034 | 339 | 38 | 99.75% | 0 | 2 | 381 |
| cdc_n2 | Probe | 14958 | 120 | 333 | 97.84% | 0 | 1 | 372 |
| cdc_n3 | Probe | 15059 | 312 | 40 | 99.74% | 1 | 353 | 36 |
| China_N | Probe | 12170 | 153 | 3088 | 79.96% | 0 | 2 | 691 |
| China_ORF1ab | Probe | 14330 | 132 | 949 | 93.84% | 1 | 0 | 367 |
| France_nCoV_IP2 | Probe | 15295 | 60 | 56 | 99.64% | 1 | 0 | 367 |
| France_nCoV_IP4 | Probe | 15326 | 53 | 32 | 99.79% | 0 | 0 | 1190 |
| HKU-N | Probe | 14947 | 129 | 335 | 97.83% | 327 | 33 | 34 |
| HKU-ORF1b-nsp14 | Probe | 15278 | 70 | 63 | 99.59% | 324 | 29 | 452 |
| Japan_NIID_2019-nCOV_N | Probe | 0 | 15079 | 332 | 97.85% | 0 | 0 | 384 |
| Japan_NIID_WH-1_F24381 | Outer | 15184 | 152 | 75 | 99.51% | 0 | 1 | 601 |
| Japan_NIID_WH-1_F501 | Outer | 15109 | 195 | 107 | 99.31% | 0 | 1 | 370 |
| Japan_NIID_WH-1_F509 | Outer | 15156 | 140 | 115 | 99.25% | 1 | 0 | 369 |
| Japan_NIID_WH-1_Seq_F24383 | Outer | 15180 | 151 | 80 | 99.48% | 0 | 0 | 606 |
| Japan_NIID_WH-1_Seq_F519 | Outer | 15004 | 232 | 175 | 98.86% | 0 | 1 | 369 |
| Japan_WuhanCoV-spk1 | Outer | 15210 | 122 | 79 | 99.49% | 1 | 1 | 594 |
| Thailand_WH-NIC_N | Probe | 15226 | 142 | 43 | 99.72% | 0 | 2 | 375 |
Updated to 17,446 COVID-19 WGS from GISAID.
Table 1. Results from PSET analysis. The five Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Along with assays from China, Hong Kong, Thailand, Japan, and France. Each assay was tested using SARS-CoV-2 as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | type | targets | PT | TP | FN | Percent True | PF | FP | TN |
|---|---|---|---|---|---|---|---|---|---|
| Noblis.12 | Probe | 2697049 | 17251 | 81 | 114 | 99.35% | 0 | 0 | 680 |
| Noblis.40 | Probe | 2697049 | 16740 | 627 | 79 | 99.55% | 0 | 1 | 1420 |
| Noblis.42 | Probe | 2697049 | 17338 | 33 | 75 | 99.57% | 0 | 2 | 1136 |
| Noblis.44 | Probe | 2697049 | 17127 | 48 | 271 | 98.45% | 0 | 1 | 1786 |
| Noblis.57 | Probe | 2697049 | 17153 | 137 | 156 | 99.11% | 0 | 1 | 853 |
| ncov_e_gene | Probe | 2697049 | 17297 | 50 | 99 | 99.43% | 361 | 15 | 53 |
| ncov_n_gene | Probe | 2697049 | 17199 | 143 | 104 | 99.40% | 0 | 340 | 72 |
| ncov_rdrp_1 | Probe | 2697049 | 0 | 17392 | 54 | 99.69% | 433 | 105 | 5967 |
| ncov_rdrp_2 | Probe | 2697049 | 2 | 17389 | 55 | 99.68% | 2 | 73 | 5916 |
| cdc_n1 | Probe | 2697049 | 16952 | 443 | 51 | 99.71% | 0 | 2 | 381 |
| cdc_n2 | Probe | 2697049 | 16939 | 137 | 370 | 97.88% | 0 | 1 | 372 |
| cdc_n3 | Probe | 2697049 | 17038 | 361 | 47 | 99.73% | 1 | 353 | 36 |
| China_N | Probe | 2697049 | 13895 | 241 | 3310 | 81.03% | 0 | 2 | 691 |
| China_ORF1ab | Probe | 2697049 | 16202 | 147 | 1097 | 93.71% | 1 | 0 | 367 |
| France_nCoV_IP2 | Probe | 2697049 | 17301 | 69 | 76 | 99.56% | 1 | 0 | 367 |
| France_nCoV_IP4 | Probe | 2697049 | 17308 | 75 | 63 | 99.64% | 0 | 0 | 1190 |
| HKU-N | Probe | 2697049 | 16938 | 145 | 363 | 97.92% | 327 | 33 | 34 |
| HKU-ORF1b-nsp14 | Probe | 2697049 | 17287 | 83 | 76 | 99.56% | 324 | 29 | 452 |
| Japan_NIID_2019-nCOV_N | Probe | 2697049 | 0 | 17076 | 370 | 97.88% | 0 | 0 | 384 |
| Japan_NIID_WH-1_F24381 | Outer | 2697049 | 17178 | 165 | 103 | 99.41% | 0 | 1 | 601 |
| Japan_NIID_WH-1_F501 | Outer | 2697049 | 16964 | 318 | 164 | 99.06% | 0 | 1 | 370 |
| Japan_NIID_WH-1_F509 | Outer | 2697049 | 17110 | 163 | 173 | 99.01% | 1 | 0 | 369 |
| Japan_NIID_WH-1_Seq_F24383 | Outer | 2697049 | 17172 | 165 | 109 | 99.38% | 0 | 0 | 606 |
| Japan_NIID_WH-1_Seq_F519 | Outer | 2697049 | 16948 | 253 | 245 | 98.60% | 0 | 1 | 369 |
| Japan_WuhanCoV-spk1 | Outer | 2697049 | 17208 | 133 | 105 | 99.40% | 1 | 1 | 594 |
| Thailand_WH-NIC_N | Probe | 2697049 | 17216 | 173 | 57 | 99.67% | 0 | 2 | 375 |
*Updated to 25,181 COVID-19 WGS from GISAID. (*small error in previous post, re-uploaded with correction)
Table 1. Results from PSET analysis. The five Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Along with assays from China, Hong Kong, Thailand, Japan, and France. Each assay was tested using SARS-CoV-2 as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | type | PT | TP | FN | Percent True | PF | FP | TN |
|---|---|---|---|---|---|---|---|---|
| Noblis.12 | Probe | 24952 | 107 | 122 | 99.52% | 0 | 0 | 680 |
| Noblis.40 | Probe | 24227 | 871 | 83 | 99.67% | 0 | 1 | 1420 |
| Noblis.42 | Probe | 25011 | 43 | 127 | 99.50% | 0 | 2 | 1136 |
| Noblis.44 | Probe | 24681 | 75 | 425 | 98.31% | 0 | 1 | 1786 |
| Noblis.57 | Probe | 24777 | 210 | 194 | 99.23% | 0 | 1 | 853 |
| ncov_e_gene | Probe | 25005 | 68 | 108 | 99.57% | 361 | 15 | 53 |
| ncov_n_gene | Probe | 24847 | 214 | 120 | 99.52% | 0 | 340 | 72 |
| ncov_rdrp_1 | Probe | 0 | 25122 | 59 | 99.77% | 433 | 105 | 5967 |
| ncov_rdrp_2 | Probe | 2 | 25119 | 60 | 99.76% | 2 | 73 | 5916 |
| cdc_n1 | Probe | 24582 | 533 | 66 | 99.74% | 0 | 2 | 381 |
| cdc_n2 | Probe | 24617 | 181 | 383 | 98.48% | 0 | 1 | 372 |
| cdc_n3 | Probe | 24675 | 399 | 107 | 99.58% | 1 | 353 | 36 |
| China_N | Probe | 18633 | 320 | 6228 | 75.27% | 0 | 2 | 691 |
| China_ORF1ab | Probe | 23801 | 150 | 1230 | 95.12% | 1 | 0 | 367 |
| France_nCoV_IP2 | Probe | 24984 | 106 | 91 | 99.64% | 1 | 0 | 367 |
| France_nCoV_IP4 | Probe | 24927 | 180 | 74 | 99.71% | 0 | 0 | 1190 |
| HKU-N | Probe | 24627 | 179 | 375 | 98.51% | 327 | 33 | 34 |
| HKU-ORF1b-nsp14 | Probe | 24942 | 145 | 94 | 99.63% | 324 | 29 | 452 |
| Japan_NIID_2019-nCOV_N | Probe | 0 | 24797 | 384 | 98.48% | 0 | 0 | 384 |
| Japan_NIID_WH-1_F24381 | Outer | 24762 | 291 | 128 | 99.49% | 0 | 1 | 601 |
| Japan_NIID_WH-1_F501 | Outer | 24614 | 388 | 179 | 99.29% | 0 | 1 | 370 |
| Japan_NIID_WH-1_F509 | Outer | 24767 | 227 | 187 | 99.26% | 1 | 0 | 369 |
| Japan_NIID_WH-1_Seq_F24383 | Outer | 24757 | 291 | 133 | 99.47% | 0 | 0 | 606 |
| Japan_NIID_WH-1_Seq_F519 | Outer | 24591 | 298 | 292 | 98.84% | 0 | 1 | 369 |
| Japan_WuhanCoV-spk1 | Outer | 24792 | 259 | 130 | 99.48% | 1 | 1 | 594 |
| Thailand_WH-NIC_N | Probe | 24878 | 232 | 71 | 99.72% | 0 | 2 | 375 |
Updated to 30,561 COVID-19 WGS from GISAID.
Table 1. Results from PSET analysis. The five Noblis assays were compared alongside the four assays from Corman and three assays from the CDC. Along with assays from China, Hong Kong, Thailand, Japan, and France. Each assay was tested using SARS-CoV-2 as the intended target. All off-target hits (TN, PF, FP) are to entries in NCBI BLAST databases (nt, gss, and env_nt).
| identifier | type | PT | TP | FN | Percent True | PF | FP | TN |
|---|---|---|---|---|---|---|---|---|
| Noblis.12 | Probe | 30281 | 139 | 141 | 99.54% | 0 | 0 | 680 |
| Noblis.40 | Probe | 29454 | 1013 | 94 | 99.69% | 0 | 1 | 1420 |
| Noblis.42 | Probe | 30352 | 60 | 149 | 99.51% | 0 | 2 | 1136 |
| Noblis.44 | Probe | 30013 | 95 | 453 | 98.52% | 0 | 1 | 1786 |
| Noblis.57 | Probe | 30033 | 267 | 261 | 99.15% | 0 | 1 | 853 |
| ncov_e_gene | Probe | 30355 | 81 | 125 | 99.59% | 361 | 15 | 53 |
| ncov_n_gene | Probe | 30153 | 266 | 142 | 99.54% | 0 | 340 | 72 |
| ncov_rdrp_1 | Probe | 0 | 30483 | 78 | 99.74% | 433 | 105 | 5967 |
| ncov_rdrp_2 | Probe | 2 | 30483 | 76 | 99.75% | 2 | 73 | 5916 |
| cdc_n1 | Probe | 29815 | 664 | 82 | 99.73% | 0 | 2 | 381 |
| cdc_n2 | Probe | 29912 | 249 | 400 | 98.69% | 0 | 1 | 372 |
| cdc_n3 | Probe | 29996 | 497 | 68 | 99.78% | 1 | 353 | 36 |
| China_N | Probe | 22004 | 375 | 8182 | 73.23% | 0 | 2 | 691 |
| China_ORF1ab | Probe | 28846 | 183 | 1532 | 94.99% | 1 | 0 | 367 |
| France_nCoV_IP2 | Probe | 30343 | 122 | 96 | 99.69% | 1 | 0 | 367 |
| France_nCoV_IP4 | Probe | 30271 | 208 | 82 | 99.73% | 0 | 0 | 1190 |
| HKU-N | Probe | 29953 | 218 | 390 | 98.72% | 327 | 33 | 34 |
| HKU-ORF1b-nsp14 | Probe | 30247 | 209 | 105 | 99.66% | 324 | 29 | 452 |
| Japan_NIID_2019-nCOV_N | Probe | 0 | 30160 | 401 | 98.69% | 0 | 0 | 384 |
| Japan_NIID_WH-1_F24381 | Outer | 30007 | 394 | 160 | 99.48% | 0 | 1 | 601 |
| Japan_NIID_WH-1_F501 | Outer | 29912 | 437 | 212 | 99.31% | 0 | 1 | 370 |
| Japan_NIID_WH-1_F509 | Outer | 30075 | 265 | 221 | 99.28% | 1 | 0 | 369 |
| Japan_NIID_WH-1_Seq_F24383 | Outer | 30002 | 391 | 168 | 99.45% | 0 | 0 | 606 |
| Japan_NIID_WH-1_Seq_F519 | Outer | 29898 | 320 | 343 | 98.88% | 0 | 1 | 369 |
| Japan_WuhanCoV-spk1 | Outer | 30039 | 352 | 170 | 99.44% | 1 | 1 | 594 |
| Thailand_WH-NIC_N | Probe | 30203 | 273 | 85 | 99.72% | 0 | 2 | 375 |